SciKit-Surgery: compact libraries for surgical navigation
Posted on 13 July 2020
SciKit-Surgery: compact libraries for surgical navigation
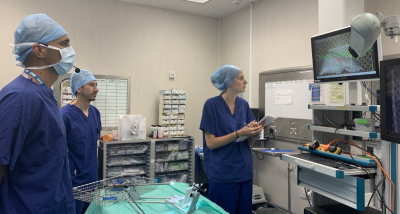 Clinicians testing the SmartLiver voice-control.
Clinicians testing the SmartLiver voice-control.
Image from paper.
By Stephen Thompson, Research Associate at WEISS, University College London.
At WEISS we are developing the SciKit-Surgery libraries to support researchers in creating better software, and to enable faster translation of research to clinical practice.
Development of the libraries is funded as part of the WEISS centre, however the libraries themselves are fully open source and we hope to see them used more widely. The SciKit-Surgery vision is to provide researchers with:
- the building blocks from which they can turn their research algorithm into a clinical application.
- the resources and training to help them turn a research algorithm into a reusable software library.
Our past experience with platforms for medical imaging and computer aided surgery (SlicerIGT, MITK, NifTK) informed our design decisions when developing SciKit-Surgery. Whilst we have been able to use these platforms to develop prototype clinical applications, we have found that it is increasingly hard to get researchers to contribute their code to the platform. The result is a central platform that requires significant software engineering resources to maintain, whilst researchers build most of their novel functionality into applications which cannot be easily reused. We identified two main reasons for this:
- choice of language - simply put most researchers find c++ (or more specifically the compilation process) very off-putting.
- loss of orthogonality within the platform. Whilst most platforms have a modular architecture in theory, in practice, as development continues cross dependencies are introduced between modules, so that eventually a programmer can no longer be confident where to implement their code or what consequences may follow a change.
Based on this, the two main design choices for SciKit-Surgery were:
- use a non-compiled language with syntax that most researchers can understand, in this case Python.
- keep each library small and independent - less than 2,000 lines of code, tested and deployed independently.
As part of SciKit-Surgery we have developed a Python template that allows researchers to develop, test, and deploy a sustainable software implementation of their algorithm. The researchers can then cherry-pick the SciKit-Surgery libraries they need to turn their algorithm into a clinical application. The ideal workflow would be something like this:
- A researcher comes up with a novel algorithm and implements it as a Python library.
- The researcher uses this implementation to do proof of concept and publish to an academic journal, including a citation to the specific version of the library used, to enable repeatable science.
- The researcher (plus some software engineering help) assembles the library into a clinical application which is used in multicentre trials to demonstrate clinical benefit.
- Results published in clinical journal and translated to a clinical product.
Obviously, as progress in science isn’t always predictable, we aim to keep the libraries flexible enough to support other workflows.
So far we have implemented around 20 libraries covering interfaces with clinical hardware, calibration, registration and visualisation. The wiki page contains a regularly updated list of libraries. Each library is hosted on GitHub and has its own issue tracking, so interested users or developers can raise issues or contribute to the code via forking the repositories and raising merge requests. The libraries are deployed to PyPi and can be easily installed with pip. The libraries were formerly known as SNAPPY, however after much debate we settled on the SciKit-Surgery name as it succinctly sums up the purpose of the libraries and is easy to search for online.
To date SciKit-Surgery has been used for two clinical applications (SmartLiver and Skull Base Navigation), along with three educational demonstrators (SciKit-SurgeryBARD, SciKit-SurgeryFRED and SnappySonic). We strongly encourage contributions to the libraries, both in code and comments or ideas for improvement. Please get in touch via the GitHub pages or by email.
Related tutorials and links:
SciKit-Surgery augmented reality tutorial.
