Enabling Reproducible Archaeology
Posted on 15 November 2021
Enabling Reproducible Archaeology
Written by Alison Clarke, with contributions from Emma Karoune.
How can we help researchers to work in a reproducible way? This blog post documents a workshop held for researchers in Durham University’s Archaeology Department, which aimed to help them make steps towards using software sustainably and working reproducibly.
I was awarded a Software Sustainability Institute (SSI) Fellowship earlier this year, with the aim of improving training in software sustainability in specific domains. The idea came from a conversation at Durham Research Methods Cafe, where Michelle De-Gruchy, a PDRA in the Archaeology department at Durham University, said that although she’d been on courses such as learning version control with git, she didn’t know how to put that into practice given her current workflows. What I’m hoping to do over the course of my fellowship is to work with researchers in one or more domains (starting with Archaeology) to determine what sort of training they need in order to be able to use software sustainably, and to produce resources that can help researchers find the right path through the many training courses that are available, to enable them to find a process that works for them.
Introducing reproducibility
As a starting point, together with Emma Karoune whose SSI fellowship aims to enhance sustainability of research in Archaeology by providing training in open science skills, I set up a workshop for researchers in the Archaeology Department, to introduce the idea of reproducibility and to try to find out more about their needs. Emma led the first part of the workshop, explaining the concept of reproducibility and how it can be applied in Archaeology (despite Archaeology being a destructive process). Emma advocated a “small steps” approach to reproducibility, to encourage researchers that they can make progress in reproducibility without having to climb the whole mountain at once! Emma described methods that can be used to enable reproducibility, and talked about data storage, and Nicholas Syrotiuk, Data Manager at Durham University, contributed to the talk with details of the pros and cons of different data repositories, including Durham’s own data repository. I then discussed reproducible analyses, with steps starting at documenting what you’re doing in a GUI tool via coding, to version control and containerisation.
The workshop then moved on to Case Studies. Emma showed some brilliant examples of how to make a project open and reproducible from data collection right through to paper publication. We then looked at some work I’d been doing alongside Michelle to automate logging and reproducibility of analyses in QGIS.
QGIS is an open-source alternative to ArcGIS, and allows users to “create, edit, visualise, analyse and publish geospatial information”. It is primarily used through its GUI but it can also be controlled via python and C++ APIs. Michelle tends to use the GUI for producing analyses, and wanted to know the best way to reproduce her work.
QGIS stores a log of processing commands that have been run, so I created a tool to parse the log and create a python script that others can run to reproduce the analysis (given the original data). The python script could then be edited if necessary, and stored in version control to allow others to change it. The tool is still a work in progress (feedback and contributions are welcome!), but we demonstrated that it could be used to record a simple analysis and replay it on a new project. My hope is that it might also encourage researchers who have completed a basic python course to see how they can use python with QGIS and start working with scripts as well as the GUI!
Finding out more about the audience
We did some live polling to get more of an idea of who our audience were and their attitudes towards reproducibility. Nearly half our audience was made up of Postdoctoral Researchers, with the remainder being Associate Professors, Assistant Professors and Postgraduates; the majority were Landscape Archaeologists, with some Environmental Archaeologists and Archaeological Scientists (Materials) present as well.
There was a wide range of software being used for analysis, including statistical software, imaging software and GIS tools, but the most-used tool was Excel; perhaps this would be a good audience for this year’s SSI Research Software Camp: Beyond the Spreadsheet?
 Fig 1: Word cloud of answers to “What software do you use for analysing your data?”
Fig 1: Word cloud of answers to “What software do you use for analysing your data?”
Most respondents scored themselves between 1 and 3 on their reproducibility journey using a scale of 1 to 5; this probably represents the scale of the challenge of creating fully reproducible research, but hopefully taking small steps will help people to make progress on that journey.
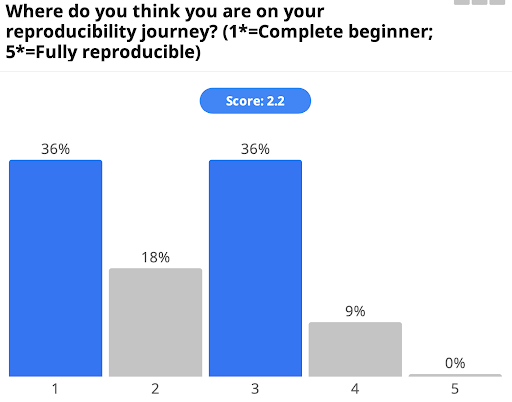
Fig 2: Graph of answers to “Where do you think you are on your reproducibility journey?”
The next question was about whether the audience thought their research group would support moving towards a reproducible approach. Two-thirds answered “Yes”, which is encouraging, but it may be worth delving deeper into the remaining one-third who answered “Maybe”, to find out what they think the barriers are.
The final question in the live poll was “What do you find most daunting about reproducibility?”. There are some interesting answers here, with the most popular being “coding”. What we need to find out next is what help people need, such as whether the internal training courses run at Durham University would fill this need or whether further resources and different courses need to be created.
 Fig 3: Word cloud of answers to “What do you find most daunting about reproducibility?”
Fig 3: Word cloud of answers to “What do you find most daunting about reproducibility?”Discussion session
Following the survey, we moved into breakout rooms to discuss data and analysis separately. In the data breakout room, attendees had questions about how data sets can be made reproducible when they contained sensitive data (e.g. high resolution geographic data); we advised instead of access upon request there should be a clear private access process held by a repository. There was also a question about intellectual property and credit for building on work done by a previous student: we advised that the university policy on ownership (of both data and interpretation) should be checked, but also that it would be most ethically responsible to preserve the original dataset in its creator’s name (and add a DOI). Further work could then cite that dataset. If researchers start to do this then it could create a virtuous circle allowing more and more data to be reused.
In the analysis breakout room, attendees asked how to sort out the wheat from the chaff when storing analyses, e.g. when a user has tried multiple parameters for an analysis and using the QGIS script I developed. We advised that with a little python knowledge and using git, the researcher could edit and update the generated script each time, rather than keeping the entire history in a single file - and that way they could go back to a previous version if necessary. We discussed data cleaning (for a particularly messy data set), which is currently being done in Excel; could that user benefit from learning regular expressions in a programming language? We also discussed using familiar GUIs to learn scripting, e.g. Deducer, which is a tool built to provide a GUI for R and designed to be a free alternative to tools such as SPSS, or DataEditR, which can be used to edit data in a way familiar to those used to using spreadsheets. These may be a less daunting place to start than learning via text files and a command prompt or even RStudio; however, it is important to ensure that any such tools record the steps taken so that the steps can be reproduced.
Next steps
How can we help these researchers on their reproducibility journey? Emma and I think we need more detail about the sorts of problems they are facing, so our next task is to hold drop-in sessions. This is to allow people to bring their issues, code and/or data, so that not only can we try to provide some next steps, but we can get a better understanding of the barriers archaeologists (at Durham University, at least!) are facing when it comes to reproducible research.
Links:


