Species distribution modellers who don’t code, want to code!
Posted on 16 January 2017
Species distribution modellers who don’t code, want to code!
 By Thomas Etherington, Senior Research Leader, Royal Botanic Gardens, Kew and Institute Fellow.
By Thomas Etherington, Senior Research Leader, Royal Botanic Gardens, Kew and Institute Fellow.
Species distribution models are a computational technique commonly used to map the likely geographic occurrence of organisms. For example, here at the Royal Botanic Gardens, Kew, we use species distribution models to help conserve plants. This conservation occurs in situ by protecting areas that models show are more likely to contain plants of interest, and also conserving plants ex situ by targeting expeditions to areas more likely to have plants from which seeds can be collected for storage in Kew’s Millennium Seed Bank.
Due to their importance in conservation, species distribution models should, like any scientific computational method, be done in an open manner so that the findings can be replicated and confirmed. Unfortunately, my experience in reading and reviewing scientific papers suggests that many scientists are still using GUI software rather than using a coding approach that enables such replication. I suspect this is probably due to a lack of computational training amongst species distribution modellers, and hence this could be something I could aim to rectify this year as part of my Software Sustainability Institute Fellowship. However, before embarking on a campaign to promote this use of sustainable software by species distribution modellers, I was curious to know if this was simply my perception, or if there was any evidence to support my own experience.
I discovered that a paper by Joppa et al. (2013) had conducted a survey of software use by species distribution modellers. The main thrust of the paper was to do with concerns about people trusting software implicitly, rather than assessing the software, with the authors recommending that software receives a greater deal of peer-review. However, the paper made the data from their survey available, and there were also some interesting data about software preferences and desires relevant to my thoughts about the prevalence of coding by species distribution modellers – see figure below.
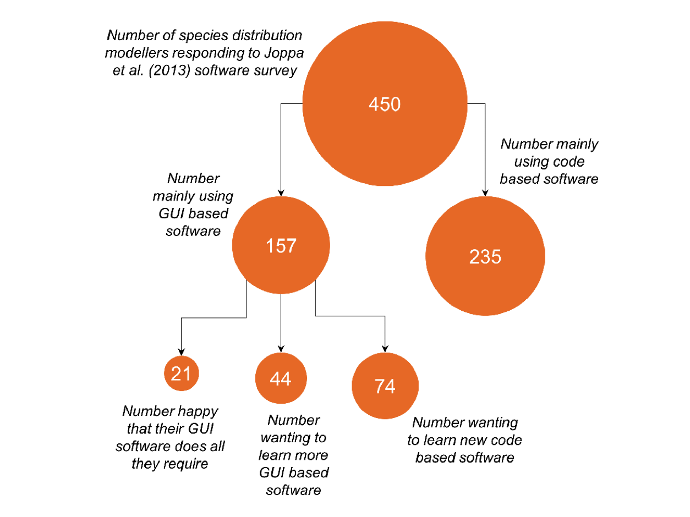
Encouragingly the majority of respondents to the survey stated a preference for using code based software, primarily R, for developing species distribution models – many of whom even identified the key coding benefits of analytical tractability and methodological transparency as main reasons for their choice! Also, although there were a large number of respondents that stated a preference for GUI software, only a small minority stated that their GUI software did all that they needed, and nearly half of these respondents identified coding as an approach that they would like to learn.
So while it would appear my perception that GUI based software is still widely used by species distribution modellers is true, the results from this survey would suggest that many of these scientists are keen to learn to code to develop future models. There is a good reason for encouraging the use of R specifically, as there are numerous R packages such as dismo, sdm, and zoon that all attempt to enable a programmatic approach to species distribution modelling. The problem here of course is that it can be a daunting process to learn computer programming, so before species distribution modellers currently using GUI software can make use of these dedicated packages some foundational R programming training is required. While organisations such as Data Carpentry provide freely available R workshop materials, including those specifically for ecology, my own experiences of learning to program have shown that trying to learn something on my own by self-teaching is much harder than learning as part of a group with the help of an instructor. Therefore, as part of my Fellowship I’ll certainly be having a think about running workshops either within an institution or as part of a conference to try and encourage species distribution modellers to adopt more open and sustainable software practices.
Photo credit: "Species Distribution Model" by Thomas Etherington is licensed under CC BY 2.0.

